CSIR - Centre for Cellular & Molecular Biology
Council of Scientific and Industrial Research
Ministry of Science & Technology, Govt. of India

J C Bose Fellow
Email: manjula@ccmb.res.in
Phone: +91-040-27192514
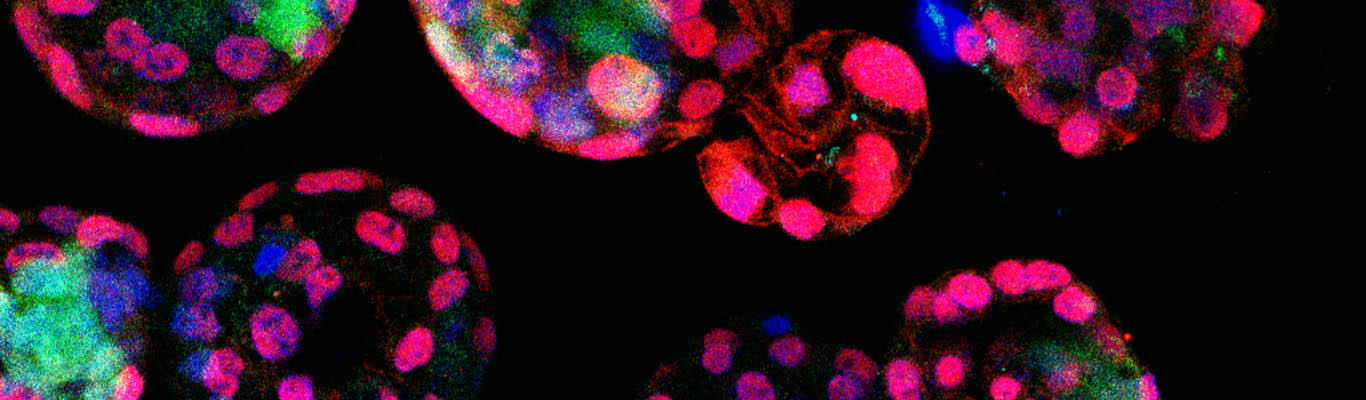
Research in our laboratory is focused towards understanding how bacteria elongate, divide, and split their cell walls during cell cycle to successfully generate two equal daughter cells. We take a multi-disciplinary approach including genetics, biochemistry, cell biology, and genomics to address these questions. We use a well-known Gram-negative rod-shaped bacterium Escherichia coli as our primary model system. Our most significant contribution in this field was identification of a rate-limiting cleavage step in the growth and synthesis of peptidoglycan, a unique constituent of bacterial cell wall. We expect that our work facilitates better understanding of fundamental aspects of bacterial physiology thereby providing novel strategies for development of alternate antimicrobial therapeutics.
Selected Publications
Rajguru V, Chatterjee S, Garde S, and Reddy M 2024. Crosslink cleaving enzymes: the smart autolysins that remodel the bacterial cell wall. Trends in Microbiology doi: 10.1016/j.tim.2023.11.004
Som, N and Reddy, M. 2023. Cross-talk between phospholipid synthesis and peptidoglycan expansion by a cell wall hydrolase. Proc. Natl. Acad. Sci USA.120 (24), e2300784120.
Bahadur R, Chodisetti PK and Reddy M. 2021. Cleavage of Braun's lipoprotein Lpp from the bacterial peptidoglycan by a paralog of L,D-transpeptidases, LdtF. Proc. Natl. Acad. Sci USA 118: 19 (2).
Chodisetti PK and Reddy M. 2019. Peptidoglycan hydrolase of an unusual cross-link specificity contributes to bacterial cell wall synthesis. Proc. Natl. Acad. Sci USA 116: 7825-7830.
Education & Experience
| P.G: | Biochemistry ; University of Hyderabad ; 1985 |
| Ph.D: | Studies of mutational mechanisms in Escherichia coli in various phases of growth: reversion analysis of mutations in the lacZ gene ; Jawaharlal Nehru University (JNU-CCMB), New Delhi ; 2001 |
|
data Found.... data Found.... data Found.... data Found.... data Found.... data Found.... data Found.... data Found.... data Found.... data Found.... data Found.... data Found.... | |
No data Found.... No data Found.... No data Found.... No data Found.... No data Found....No data Found.... No data Found.... No data Found.... No data Found.... No data Found....
| Title | Journal | Year |
|---|---|---|
| Glycan strand cleavage by a lytic transglycosylase, MltD contributes to the expansion of peptidoglycan in Escherichia coli. | PLoS genetics | 2024 |
| Crosslink cleaving enzymes: the smart autolysins that remodel the bacterial cell wall. | Trends in Microbiology | 2023 |
| Cross-talk between phospholipid synthesis and peptidoglycan expansion by a cell wall hydrolase | Proceedings of the National Academy of Sciences | 2023 |
| Peptidoglycan Remodeling by an L,D-Transpeptidase, LdtD during Cold Shock in Escherichia coli | Journal of Bacteriology | 2022 |
| Modulation of RecFORQ- and RecA-mediated homologous recombination in Escherichia coli by isoforms of translation initiation factor IF2 | Journal of Bacteriology | 2022 |
| Structural basis for the peptidoglycan editing activity of YfiH | mBio | 2022 |
| A LytM-factor, ActS functions in two distinctive peptidoglycan hydrolytic pathways in Escherichia coli | Frontiers in Microbiology | 2022 |
| Cleavage of Braun`s lipoprotein Lpp from the bacterial peptidoglycan by a paralog of L,D-transpeptidases, LdtF | PNAS | 2021 |
| Peptidoglycan: structure, synthesis and regulation | EcoSal Plus | 2021 |
| Structural basis for the differential regulatory roles of the PDZ domain in C-terminal processing proteases. | mBio | 2019 |
| Peptidoglycan hydrolase of an unusual cross-link specificity contributes to bacterial cell wall synthesis. | Proc Natl Acad Sci. USA | 2019 |
| A synergistic role for two predicted inner membrane proteins of Escherichia coli in cell envelope integrity. | Molecular Microbiology | 2019 |
| Structural basis of adaptor-mediated protein degradation by the tail-specific PDZ-protease Prc. | Nature Communications | 2017 |
| Identification of YfiH (PgeF) as a factor contributing to the maintenance of bacterial peptidoglycan composition. | Molecular Microbiology | 2017 |
| Determination of Keto-deoxy-d-manno-8-octanoic acid (KDO) from Lipopolysaccharide of Escherichia coli. | Bio-protocol | 2015 |
| Regulated proteolysis of a crosslink-specific peptidoglycan hydrolase contributes to bacterial morphogenesis. | Proc. Natl. Acad. Sci USA | 2015 |
| yciM is an essential gene required for regulation of lipopolysaccharide synthesis in Escherichia coli. | Molecular Microbiology | 2014 |
| Three redundant murein endopeptidases catalyse an essential cleavage step in peptidoglycan synthesis of Escherichia coli K12. | Molecular Microbiology | 2012 |
| SufI (FtsP) is a cell division protein required for the growth of Escherichia coli during oxidative stress. | Journal of Bacteriology | 2007 |
| Role of FtsEX in cell division of Escherichia coli: viability of ftsEX mutants is dependent on functional SufI or high osmotic strength. | Journal of Bacteriology | 2007 |
| Intersubunit signaling in RecBCD enzyme, a complex protein machine regulated by Chi hotspots. | Genes & Development | 2007 |
| A positive selection system for identification of recombinants using alpha-complementation plasmids. | BioTechniques | 2004 |
| Distinct signatures for mutator sensitivity of lacZ reversions, and for spectrum of lacI/lacO forward mutations on the chromosome of nondividing Escherichia coli. | Genetics | 2004 |
| IS186 insertion at a hot spot in the lon promoter as a basis for Lon protease deficiency of Escherichia coli B: Identification of a consensus target sequence for IS186 transposition. | Journal of Bacteriology | 2001 |
| lon incompatibility associated with mutations causing SOS induction: Null uvrD alleles induce an SOS response in Escherichia coli. | Journal of Bacteriology | 2000 |
| Characterization of the uup locus and its role in transposon excisions and tandem repeat deletions in Escherichia coli. | Journal of Bacteriology | 2000 |
| A genetic strategy to demonstrate the occurrence of spontaneous mutations in nondividing cells within colonies of Escherichia coli. | Genetics | 1997 |
| Identification and characterization of ssb and uup mutants with increased frequency of precise excision of transposon Tn10 derivatives: nucleotide sequence of uup in Escherichia coli. | Journal of Bacteriology | 1997 |


2514
manjula@ccmb.res.in

2519
krishnasree@ccmb.res.in

2521
shambhavigarde@ccmb.res.in
2521
surajkumar@ccmb.res.in


2519
krishnachaitanya.n@ccmb.res.in

2521
s.chatterjee@ccmb.res.in
2521
vishnu.vachana@ccmb.res.in

2521
vaidehi.rajguru@ccmb.res.in

2521
debnita@ccmb.res.in

2519

2521
harikrishnans@ccmb.res.in

